FRI Microbial Biofilms Research Stream
Meeting Microbes Where They’re At
In this stream, research focuses on microbial biofilms, which are living, microbial communities that develop and persist at interfaces in aqueous environments. Biofilms are composed of cells immobilized at a surface or interface that is embedded in an organic polymer matrix of microbial origin. Biofilm cells differ from their planktonic, free-swimming counterparts in the genes and proteins they express, resulting in distinct phenotypes that include altered resistance to antimicrobial agents, antibiotics, and host immune systems. Biofilms occur naturally in open environments, such as in Yellowstone National Park’s hot springs, and in closed environments. In fact, 99% of all microbial activity in the open environment is in multi-species biofilms. Biofilms can be beneficial, including use in waste-water and toxic sites, where bacteria are selected for ability to utilize or detoxify contaminants and then introduced to contaminated sites. Biofilms also occur in closed systems. For example, biofilms can occur as contaminants in industrial systems and as pathogens in organisms.
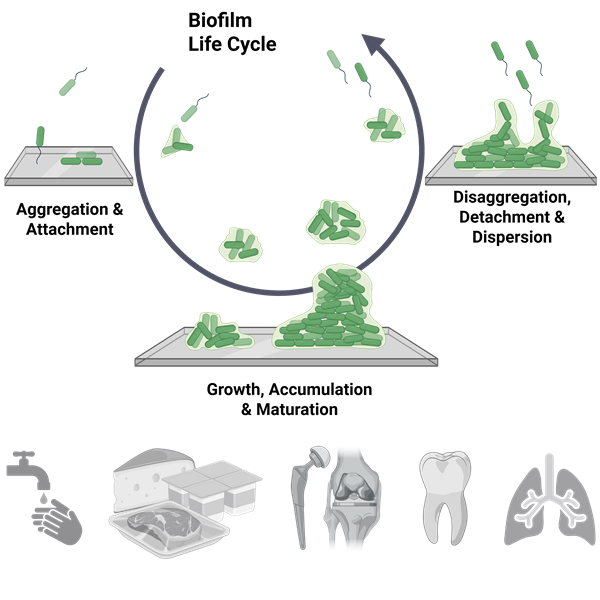
The stream works hand in hand with the core faculty members of the Binghamton Biofilm Research Center (BBRC) to investigate the basic biology of microbial biofilms in ways that inform a better understanding of how to address the problems that these biofilms can create and how to use biofilms to solve problems. Thus, FRI students can easily comprehend the relevance of basic research on biofilms and visualize the importance of collaboration in tackling real world problems.
The Microbial Biofilms in Human Health research stream utilizes a strong interdisciplinary
and collaborative approach to microbial biofilms research. In addition to the foundational
background of microbiology, students learn basic skills and techniques in cell and
molecular biology, biochemistry, and microbial ecology. Successful research in microbial
biofilms requires collaborations with faculty both inside and outside of the biofilm-research
group to bring together the specific expertise required to tackle each unique research
question.
Research Themes
Bacterial Genetics and Cell Biology
Microbial Interactions
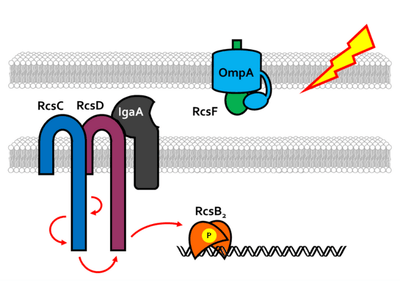
Stress Response and Signaling
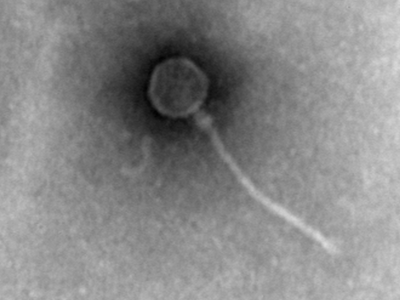
Phage-Biofilm Interactions

Antibiotics and Bacterial Toxins

Pathogenesis and Clinical Relevance
Research Educator
Bacterial cell biology Stress response and signaling systems Genetic screens Live imaging Phage and bacterial interactions
Trevor Cross
Biofilms, Research Assistant Professor
Research Interests
Dr. Trevor Cross is the FRI Research Educator for the Microbial Biofilms research stream. With broad research interests in bacterial and microbial cell biology, Dr. Cross aims to guide students in exploring a myriad of exciting topics in microbiology that contribute novel findings to the field of microbiology. The research projects possible in the Microbial Biofilms stream involve looking at interactions among different bacterial species (particularly those associated with teeth and dental health), how phage impacts bacterial biofilms, antibiotic stress responses, cellular signaling, and certainly many more exciting topics! Dr. Cross himself was a student in the SEA-PHAGES program where he was able to participate in research from his first year in college, so he understands first-hand the benefits, challenges, rigor, and joy of being in a research-intensive program like FRI early on in college and hopes he can carry on the tradition of outstanding mentorship and scholarship he experienced.
Research Techniques
Lab Work
Skills
- Pipetting
- Dilution
- Media and Solution prep
Techniques

- Aseptic technique
- Bacteria cell culture
- Molecular biology
Biofilm Models

- Well-plate growth
- Continuous-culture growth
Analysis
Quantification and Visualization

- Spectrophotometry
- Microscopy
Basic Statistics

- Excel
- Graphpad prism
Biofilm Phenotype
- Bacterial attachment
- Antibiotic susceptibility
- Dispersion and detachment
Research Projects
-
Cohort 11 (2024-2025)
- Biofilm structure and development in Candida albicans and Aspergillus fumigatus: Impacts on antifungal efficacy
- Determining the impacts of artificial sugars on the gut microbiome
- Environmental impacts on the gut microbiome triggering pediatric allergies
- Immobilized antimicrobial peptide coating as prevention against bacterial biofilm formation on urinary catheters
- The application of cranberry extract to treat biofilm-associated oral infections

-
Cohort 10 (2023-2024)
- Developing a protocol to analyze the potential of OMV-induced dispersion of Pseudomonas aeruginosa biofilms
- Genes behind the scenes: Enterococcus faecalis biofilm dispersion
- Investigating the effect of PqsA and OprF on reversible and irreversible attachment of Pseudmonas aeruginosa
- The effectiveness of household cleaners to reduce Fusarium spp. biofilms found in water systems
- Utilizing garlic to prevent Escherichia coli attachment on food industrial surfaces
-
Cohort 9 (2022-2023)
- Investigating the effect of the PAI-1 protein on the development of recurrent UTIs and potential treatment methods using antibiofilm enzymes
- Optimization of a treatment for Pseudomonas aeruginosa biofilms using antibiotics and α-amylase
- The application and proteomic analysis of outer membrane vesicles to disperse biofilms
- The effect of passive and active dispersion on virulence
- α-Amylase and tobramycin-loaded nanoparticles for the treatment of Pseudomonas aeruginosa biofilms
-
Cohort 8 (2021-2022)
- Characterization of motility and susceptibility of ɑ-amylase dispersed cells in Pseudomonas aeruginosa
- Exploring the impact of a diabetic environment on Pseudomonas aeruginosa biofilm dispersion induced by ɑ-amylase
- Implications of relaxed selection towards antibiotic resistance in Enterococcus faecium
- Investigating capacity of membrane vesicles to induce biofilm dispersion
- Optimizing a continuous-flow model to support biofilm growth of vancomycin-resistant Enterococcus

-
Cohort 7 (2020-2021)
- Comparing inhibitory ability of outer membrane vesicles from Pseudomonas aeruginosa at various stages of growth
- Inhibition of biofilm development through peptide Interference
- Inhibiting biofilm-specific tolerance via peptide interference
- Investigating the relationship of cis-2-decenoic acid and outer membrane vesicles in Pseudomonas aeruginosa
- Investigating the Pseudomonas quniolone signal as a primary inducer of Pseudomonas aeruginosa biofilm dispersion

-
Cohort 6 (2019-2020)
- Analysis of Pseudomonas quinolone signal and outer membrane vesicle production during varied biofilm dispersion responses
- Assessing predation abilities of outer membrane vesicles from Pseudomonas aeruginosa planktonic, biofilm, and dispersed cells
- Degradative enzyme abilities of outer membrane vesicles from Pseudomonas aeruginosa biofilms
- Role of Pseudomonas quinolone signaling and outer membrane vesicle production in P. aeruginosa biofilm dispersion
- The physiological role of outer membrane vesicles in Pseudomonas aeruginosa biofilms
-
Cohort 5 (2018-2019)
- Comparing microbiomes between acute and carrier group A Streptococcus pharyngitis
- Discerning the Role of Biofilm in Atherosclerotic Plaques
- Pathways Contributing to Dispersion-Related Phenotypes in Pseudomonas aeruginosa
- Variation in outer membrane vesicle proteomes from Pseudomonas aeruginosa biofilms
- Virulence of Outer Membrane Vesicles Produced by Pseudomonas aeruginosa during Biofilm Dispersion

-
Cohort 4 (2017-2018)
- Development of a Novel Quantification Method for Cyclic-di-GMP Using Bis(p-nitrophenyl) Phosphate
- Development of a Novel Quantification Method for Cyclic-di-GMP Using cMANT
- Pseudomonas Quinolone Signal Quantification During Pseudomonas aeruginosa Biofilm Development
- Quantifying Outer Membrane Vesicles During Pseudomonas aeruginosa Biofilm Development
- Role of the Type VI Secretion System on Virulence and Pathogenesis in Biofilm Dispersion

-
Cohort 3 (2016-2017)
- Pseudomonas aeruginosa cis-2-decenoic acid induced biofilm dispersion: regulation of pseduomonas quinolone signal and outer membrane vesicle production
- Gene conservation in biofilm formation genes of Pseudomonas aeruginosa and Escherichia coli
- The effect of cis-2-decenoic acid and dispersion on gene expression, pseudomonas quinolone signal, and outer membrane vesicles in Pseudomonas aeruginosa
- The role of Pseudomonas aeruginosa Las and Rhl quorum sensing networks on development and persistence of dual-species biofilms with Staphylococcus aureus
- Creating a Pseudomonas aeruginosa reporter strain to indicate native biofilm dispersion

-
Cohort 2 (2015-2016)
- The inhibition of Staphylococcus aureus wild type biofilm growth by Pseudomonas aeruginosa wild type and lasI mutant biofilms
- The effect of cis-2-decenoic acid, antibiotics, and the ppx gene on Pseudomonas aeruginosa persister cells
- Cyclase expression, sagS, and its relation to biofilm development
- Outer membrane vesicle contribution to antibiotic tolerance in Pseudomonas aeruginosa
-
Cohort 1 (2014-2015)
- Role of pyruvate in biofilm formation and antibiotic tolerance
- Role of ppx in reversion from persister state with use of cis-2-decenoic acid
- Effect of adding cis-2-decenoic acid, dispersion inducer, on antibiotic efficacy against S. aureus biofilm bacteria
- Formation and tolerance of PAO1 persister cells to antibiotic treatment
- Outer membrane vesicles and antibiotic resistance

Research Stream Collaborators

Research Interests

Research Interests

Caitlin Light
Assistant Director, Researcher, Microbial Biofilms in Human Health
Research Interests

Research Interests

Research Interests

Jeffrey W. Schertzer
Associate Professor; Biochemistry Program Director
Research Interests
No profile found.